- 4 Level Data Slicer As Seen On Tv
- Data Slicers Excel
- 4 Level Data Slicer Attachment
- 4 Level Data Slicer Manual
The Data Slicer provides an interface which allows users to get subsections of either VCF (VCFtools) or BAM (SAMtools) files based on genomic coordinates.
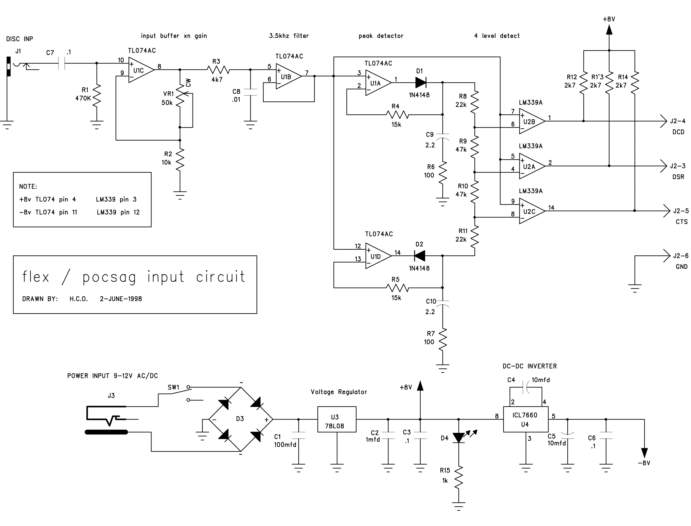
Version 4.0 revision 29402 built 2020-10-05 Preview Release Unstable! Version 4.13.0 revision 29505 built 2020-12-06: version 4.13.0 revision 29505 built 2020-12-06: version 4. Create a slicer visual for your report and then select a date value for the Field value. Select the slicer on your canvas and then the carat in the upper-right corner of the slicer visual. If the visual has date data, the menu will display the option for Relative. For the relative date slicer, select Relative. To monitor 4 level FLEX, ERMES and other data signals (POCSAG, 2 level FLEX), this 4 level interface was developed. From 2003 it can be purchased from this site. Recently, the circuit board has been redesigned, resulting in a more compact board (about 15% smaller). 4 level FSK INTERFACE 2.0 - YouTube. Block diagram of the data-slicer blocks in the MAX1473, including external components. The Fundamental Data-Slicing Circuit Figure 4 shows the simplest data-slicing circuit. The output of the data filter, DFO, goes to the positive pin of the data slicer comparator, DSP, and also forms the slicing threshold voltage at DSN by passing.
In the Ensembl GRCh37 browser, you can access the Data Slicer from the tools link in the menu bar at the top of every page.
The input interface is shown below:
The tool allows you to pick which phase of the 1000 Genomes Project you want to get data from. If you have a publicly visible VCF file and corresponding tabix index (.tbi) in the same folder, you could get data from these by selecting “Provide file URLs”.
You can select filtering by either individuals or populations. Select one to get extra options. To select multiple populations or individuals please hold the ctrl key (on windows/linux) or the cmd key (macs).
If you select BAM files, you must provide a publicly visible URL for your BAM file which must be accompanied by BAM index (.bai) of the same name. Please note that this service will only work for other BAM files over http.
After clicking “Run” the system produces your final file.
This presents a link to download your file slice which should contain both the location you requested and the name of the original in its name. You also get a summary of the first few lines of the file in the window so you can check it has produced something close to expectation.
There is also documentation for our Variation Pattern finder which looks at shared patterns of variation between individuals from VCF files.
If you have any questions about this system please email [email protected]
Home < Slicer4:Download Data- 4Development status
- This is a first generation mock-up for a database display/download schema for Slicer
- The NA-MIC download data can be used for prototyping.
- thumbnails should be extracted for now from the center slice of the volume. Mechanism to replace manually. Down the road, we will use the thumbnail from the mrml file.
- Annotation should be extracted from the DICOM headers, if available. We will need a painless way to edit the annotation (with wiki style history). Once the slicer annotation module is working (summer 2011), we can extract information from there.
- Pack as many thumbnails as possible horizontally (based on width of browser window and font size)
- Only list one or two lines of text, the rest pops up on rollover.
- Pack densly, use space sparingly
| Brain: Multi-modality (sMRI, DTI, fMRI) from Schizophrenia Study There are 20 cases: ten are Normal Controls and ten are Schizophrenic. Each case includes a weighted T1 scan, a weighted T2 scan, an fMRI scan, a DTI volume, the DWI with 51 directions, and several masks and labelmaps. Available from Harvard. |
| Brain: 2-4 Year Old from Autism Study Data for 2 autistic children and 2 normal controls (male, female) scanned at 2 years with follow up at 4 years from a 1.5T Siemens scanner. Files include structural data, tissue segmentation label map and subcortical structures segmentation. Available from UNC. |
| Brain: White Matter Lesions for Lupus Study Data for 5 cases of Lupus White Matter Lesion patients. The data is co-registered. Each case contains: T1-weighted, T2-weighted, FLAIR, and masks for brain and lesions. Available from MIND. |
- Case 1, study 1
- Lupus001
- last update on 2010-04-12 14:32:11 (13.4 MB in 8 files)
- Brain MRI study of a subject with slight Lupus Erythematodes.
- Download this study: Just the data, data with a mrml file for Slicer 3.6
- lupus001_T1.nii.gz
128x256x256 voxels
registered and bias corrected T1 series
5.7MB - uploaded on 2010-08-10 19:35:20 by XXX
Download, download with mrml - lupus002_FLAIR_reg+bias.nii.gz
128x256x256 voxels
registered and bias corrected Flair acquisition
4.86MB - uploaded on 2010-08-10 19:35:20 by xxxx
Download, download with mrml - lupus003_T2_reg+bias.nii.gz
192x256x256 voxels
registered and bias corrected T2 wieghted image
6.34MB - uploaded on 2010-08-10 19:35:20 by xxxx
Download, download with mrml
4 Level Data Slicer As Seen On Tv
- Case 1, study 2
- Lupus002
- last update on 2010-04-12 14:32:11 (13.4 MB in 8 files)
- Brain MRI study of a subject with severe Lupus Erythematodes.
- Download this study: Just the data, data with a mrml file for Slicer 3.6
- lupus001_T1.nii.gz
128x256x256 voxels
registered and bias corrected T1 series
5.7MB - uploaded on 2010-08-10 19:35:20 by XXX
Download, download with mrml - lupus002_FLAIR_reg+bias.nii.gz
128x256x256 voxels
registered and bias corrected Flair acquisition
4.86MB - uploaded on 2010-08-10 19:35:20 by xxxx
Download, download with mrml - lupus003_T2_reg+bias.nii.gz
192x256x256 voxels
registered and bias corrected T2 wieghted image
6.34MB - uploaded on 2010-08-10 19:35:20 by xxxx
Download, download with mrml
- A. Support for local customization
1. Add SQL schemas to link a resource to a template. Done
2. Add support for administrators to create/edit links between resources templates. Done
3. Modify the core MIDAS to support templates at the resource level. Done (communities, collections)
- B. Specific Customization for NAMIC datasets
1. Display of the download page at the “collection” level according to the layoutthe wiki.
-Layout : Done
-Download All: Done
-Download All with mrml: Done
-Download single bitstream: Done
-Download single bitstream with mrml: Done
-Download single resource: Done
-Download single resource with mrml: Done
-Show Items information: Done
-Show Bitstreams Metadata: Done
2. Support for manually replacing a thumbnails Done
3. Implementation of javascript support to extend description on mouse over. Done (community page)
- C. Testing phase Done
- D. Upgrade of the MIDAS server Done
Demo
Screenshots
- Community level
- Collection level
- Manage thumbnails
How to link a template to a resource
First, go resource edit page:
Then select the template you want to use:
You can recursively apply the template. (All the children of the resource will use the selected template)
How to upload new files
First, go a collection page and create an item:
Click on 'Upload Files':
Select the files you want to upload (you can select multiple files)
If Midas warns you that a file is too big, use the 'large files' tool:
When your files are uploaded, you can ask midas to create resources. If you do that, Midas will analyze the selected files and try to find the metadata:
Data Slicers Excel
How to change a thumbnail
First, go the an item page:
4 Level Data Slicer Attachment
Then go a the menu and click on 'thumbnails':
Select the thumbnail you want to change:
Upload a new image (the application will resize the image)
If you like the new thumbnail, click on 'Set thumbnail' to finalize the modification
4 Level Data Slicer Manual
Retrieved from 'https://www.slicer.org/w/index.php?title=Slicer4:Download_Data&oldid=20360'
